by Erick Figueroa Ildefonso
In human language, different words or expressions can have the same meaning. In genetics, we can find a similar situation. We can think of DNA as a four-letter alphabet, composed of A (Adenine), C (Cytosine), G (Guanine), and T (Thymine), with T replaced by U (Uracil) in RNA. DNA can store instructions for life; these instructions are copied into messenger RNA (mRNA) for eventual translation into a protein.
Now, the message in the RNA is composed of a short three-letter word named “codon”, and each codon is translated to an amino acid (e.g., AUG for methionine). There are 20 amino acids, and if we do the math, our four letters (A, C, G, T/U) can be combined into 64 three-letter codons. Of these, three codons serve as stop signals during translation (UGA, UAG, UAA), leaving us with 61 codons for 20 amino acids.
Thus, the instructions encoded in DNA and transcribed into mRNA are encoded using this 61- word code for the 20 amino acids and three stop codons that will be translated later into the amino acid sequence in the instructions. However, due to the lack of correspondence between the number of codons and the number of amino acids, we can say that the genetic code is redundant (i.e, there is degeneracy of the genetic code). The synonymous codons code the same amino acid, as is shown in the table below (Figure 1).
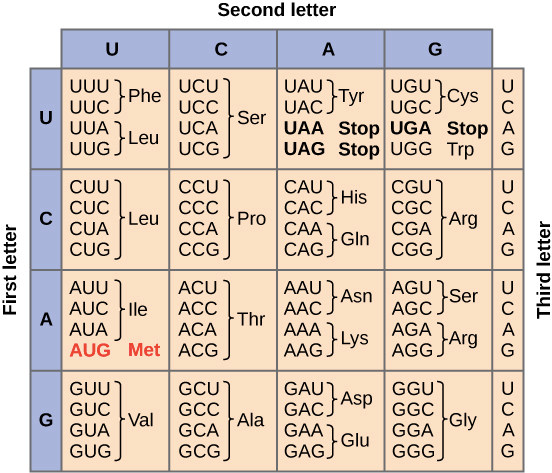
Figure 1. Genetic code table.
Image source: “The genetic code,” by OpenStax College, Biology (CC BY 3.0).
When we find that for a single amino acid certain codons, of the possible options, are preferentially or non-randomly used, we can say that there is a bias in the codon usage (Figure 2). This codon usage bias can occur as a consequence of silent mutations, which don’t change the message (i.e, don’t lead to a change in the amino acid sequence) and happen by chance. But sometimes a codon is preferentially used because its presence in a gene may optimise the translation process and is favored by natural selection.
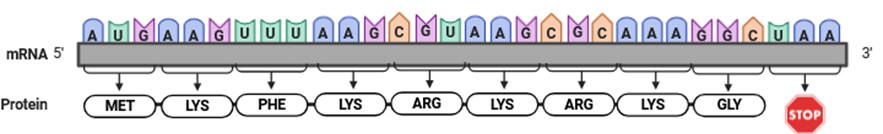
Figure 2: mRNA molecule with a more frequent use of the AAG codon for the lysine amino acid.
Image generated by the author using Biorender.
Codon preference in an mRNA is correlated with the abundance of the transfer RNAs (tRNA) carrying the anti-codon that is complementary and carries the corresponding amino acid. tRNA abundance reflects both evolutionary adaptation to typical conditions and, in some cases, regulatory responses to environmental changes. As a result, codon usage can evolve to match the pressure an organism experiences in its niche. For example, research across different yeast species has found that the enzyme required for galactose digestion shows codon usage bias, with optimization in species more frequently exposed to this sugar.
So, even though a silent mutation does not affect the amino acid sequence of a protein, it can still have consequences at other levels of an organism’s biology, including translation and gene expression.
