by Wendy Sun
Fun Rating: 3/5

Difficulty Rating: 3/5

What is the general purpose?
Complementary RNA sequencing, or RNA-seq, allows scientists to read the order of the bases (adenine, uracil, guanine, and cytosine) in an RNA molecule from a cell or tissue. Compared to direct RNA-seq, which sequences single-stranded RNA directly, complementary RNA sequencing achieves higher accuracy by stabilizing single-stranded RNA through complementary DNA pairing.
Why do we use it?
Different cells activate different sets of genes. If you think about the cells that make up your skin versus the cells that create your hair, they produce various types of proteins and thus need to use different combinations of genes. Most of these genes are controlled by information carried in RNAs. RNAs act as ‘messages’ that tell the cell which proteins to make. By reading many RNAs at once, RNA-seq tells us which genes are ‘on’ at one moment in time, thus hinting at the expression level of the RNA of interest.
One example is the use of RNA-seq in understanding how cancer cells differ from normal cells. Researchers can isolate RNA from a tumor and from matching healthy tissue, perform RNA-seq on both samples, and compare which genes are uniquely expressed in the cancer. This reveals which genes are ‘on’ (or active) at one moment in time and abnormally active in cancer cells.
How does it work?
You can think of RNA-seq as giving a summary of text messages being sent inside a cell. The steps below are how we collect and read those messages. These steps are visualized in Figure 1.
- RNA extraction: opening the cells
- This method has been previously described on our blog! In short, the purpose of this step is to open the cell, release and collect all the RNA, like opening a passion fruit and only keeping the seeds.
- RNA selection: choosing which messages to keep
- There are different types of RNA inside a cell (rRNA, tRNA, mRNA, etc). This process uses specific tools, such as gel electrophoresis, to isolate the RNA of interest.
- cDNA synthesis: making a stable copy
- RNA is single-stranded and less stable than DNA. To increase the stability of RNA, we can use specific enzymes to read RNA and build a complementary matching DNA strand, called complementary DNA (or cDNA), making a stable copy of RNA for future experiments!
- Library preparation: getting the message ready for reading
- This step allows scientists to collect all cDNAs in short fragments, making it easier for the Next-Generation Sequencing instrument to read through.
- Sequencing: reading through the letters
- The cDNA library (stands of cDNA) is inserted into an instrument (usually through next-generation sequencing), and the machine outputs the order of the RNA bases – the As, Ts, Cs, and Gs nucleotide bases we are familiar with.
- Analysis: understanding messages carried in the sequencing result
- After sequencing, a data analysis specialist will first check that the raw data pass key quality control checks to ensure the results are reliable. Next, the sequencing reads are aligned to a known RNA database to identify which RNAs are enriched in the sample. From this data, scientists can see which RNAs are more or less abundant in the sample and infer what “messages” the cells were sending at the time, such as which pathways were active or how the cells were responding to a treatment.
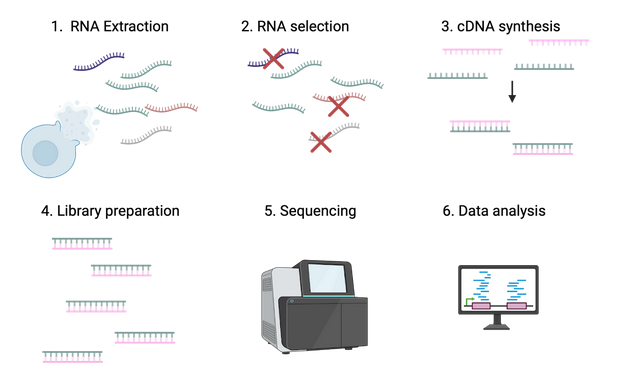
Figure 1: Schematic of RNA sequencing steps. Image created by author using BioRender.
