by RP Pornmongkolsuk
Fun Rating: 4/5

Difficulty Rating: 4/5

What is the general purpose?
Not all scientific research is done on a lab bench! Geneticists use simulations to study evolutionary processes that might be impossible to replicate experimentally or to confirm a population pattern they observed.
Why do we use it?
In population genetics, scientists observe how populations or species change their genetic makeup over time—often longer than one’s lifetime to practically design an experiment for. So, they can use powerful evolutionary simulation tools like SLiM or msprime to create even thousands of evolutionary scenarios and then use statistical calculations to infer which scenarios likely happened that led to the observed population pattern.
How does it work?
Over the past century, geneticists have developed theories of how a genetic makeup of a population can change over time. For example, how often do mutations occur and how often are they beneficial to the organisms that carry them? Or how much does the population size impact the influence natural selection has on a population? With computers as powerful as we have now, geneticists can now gather all the theoretical knowledge and implement how evolutionary processes may have worked in a program—just like how you are able to tell a calculator to perform various operations in order!
Consider SLiM, one of the most powerful simulation tools available, it can run through thousands of years worth of evolutionary processes of a population in under an hour!! A computer simulation may sound abstract, so Figure 1 shows what a simple simulation on SLiM looks like. This is a SLiMgui or SLiM graphical user interface which has the ability to show us exactly what is happening in the simulation in real-time.
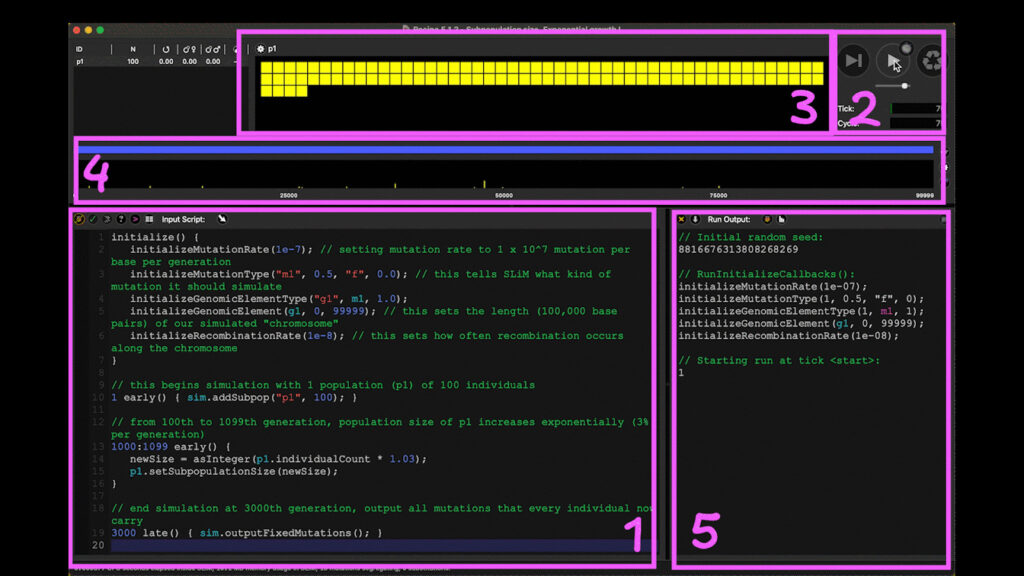
Figure 1. Window of SLiMgui simulating a simple evolutionary model.
Like many other programs, SLiM requires the input to be scripted in a particular programming language. But worry not, SLiM comes with a manual and many simple “recipes” that you can adapt from. For example, Box 1 of Figure 1 contains the script adapted from a preset recipe of an evolutionary model involving a neutral evolution of a population starting from 100 individuals growing exponentially. Neutral evolution means that natural selection does not occur in this simulation, and that all the changes are happening due to chance alone.
The script in Box 1 specifies the scenario we want SLiM to simulate, starting from setting the rate of mutation and initial population size. Then, we can specify how many generations we want SLiM to simulate and how the population may change in size at a particular point in time.
You simply start the simulation by clicking the play button in Box 2, and SLiMgui will visualize the changes in real-time. One of the changes in the recipe is population size increasing. Each yellow square in Box 3 represents one individual in the population which grew exponentially during the simulation. SLiM also generates random mutations along the chromosome. Box 4 shows positions of those mutations on the chromosome, and the yellow vertical bars represent the proportions of individuals in the population that carry a mutation at each particular position.
Finally, we ask SLiM to print some output of the simulation, in this case, all the mutations that had arisen and that, by the end, every individual in the population carries just by random chance. At the end of the simulation, SLiM outputs the list of those mutations in Box 5. These are essentially the mutations of which their yellow bars in Box 4 hits the top of the box!
Because we cannot go back in time to observe population patterns over time in the past, we can only rely on observations that we can make now. Geneticists can compare this simulated result and its statistical measures and see if this evolutionary model could explain the patterns we observed.
