by Erin Dickert
Fun Rating: 4/5

Difficulty Rating: 4/5

What is the general purpose?
Amplicon sequencing is a molecular biology technique that uses polymerase chain reaction (PCR) to amplify a short, specific sequence of DNA called an “amplicon,” followed by detailed sequencing to identify genetic variation within the amplicon.
Why do we use it?
Amplicon sequencing is a versatile next-generation sequencing technique that offers many advantages for biomedical research. Broadly, sequencing is a technique that allows scientists to determine the order of the four bases that make up DNA, and it is helpful in disease diagnosis, genetic testing, and biomedical research. Compared to whole genome sequencing, which is another next-generation sequencing technique that covers the entire genome, this method is a highly targeted sequencing approach that is specific, relatively inexpensive, and has a fast turnaround time in comparison. Researchers might use this technique when they are interested in mutations at particular sites in the genome, such as in clinical diagnostic applications. On a basic sciences level, researchers can use amplicon sequencing to identify mutations generated by CRISPR editing, allowing them to accurately ensure the modifications made to the DNA. In a more translational context, amplicon sequencing is used to identify mutations that are specific to tumors, allowing researchers to understand the genetic basis of cancers. Additionally, researchers can analyze evolving variants of viruses like SARS-CoV-2, where they can trace transmission patterns and identify new variants.
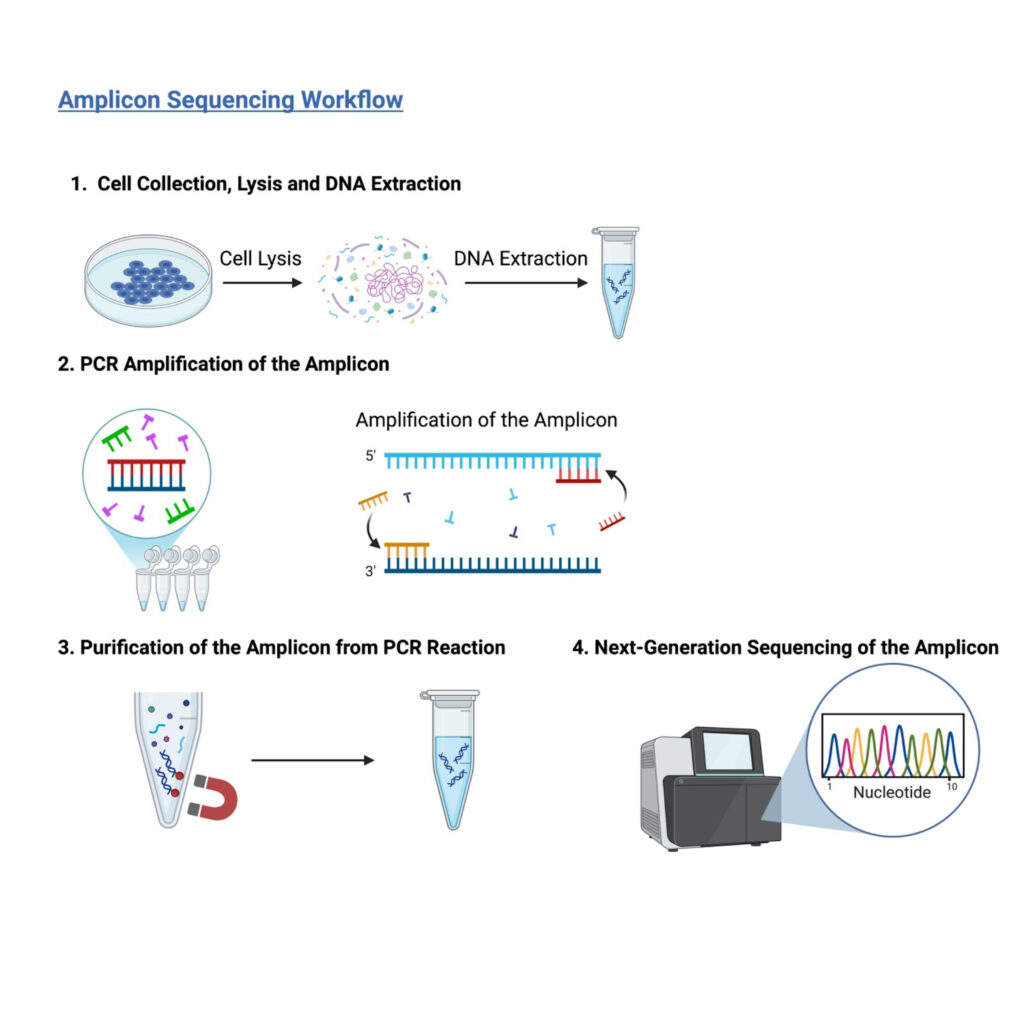
Figure 1: Steps of Amplicon PCR. The DNA is removed from the cells, amplified using PCR, purified, and sequenced using next-generation sequencing techniques. (Image created by author using BioRender)
How does it work?
1. To perform amplicon sequencing, the cells of the tissue of interest must first be collected and the cells must be lysed, or broken down, to release their contents (Figure 1.1). One could use either cells grown within a petri dish or patient samples. The DNA must then be removed from the cells, and this can occur using commercial kits that bind DNA using a silica membrane or using chemicals like phenol to separate the DNA from the proteins. If there is usable DNA, it should work for amplicon sequencing!
2. After extracting the DNA, PCR can be used to amplify the DNA sequence of interest from all the genomic DNA (Figure 1.2). As a refresher on PCR, you can read this post from the NIH! To only amplify the DNA sequence of interest, a researcher will use short pieces of DNA called primers. These short DNA sequences flank the DNA sequence of interest and are complementary to the DNA. This means that they will perfectly match up with a portion of the DNA sequence on either side of the sequence of interest. These act as starting points for the DNA synthesis step in PCR and ensure that only the DNA of interest is amplified.
3. After the DNA is amplified and we have our amplicon of interest, the amplicon must be purified to remove it from all the leftover genomic DNA, proteins, and salts left over from the PCR reaction (Figure 1.3). This can be done in several ways. Two standard methods are using a column containing a silica membrane to which the amplicon will bind, allowing the proteins and salts to be washed off, or magnetic beads, which also bind the DNA and allow the other molecules to be washed off.
4. After these first three steps, there needs to be a way to quantify, or measure, the product (Figure 1.4). This quantification is often performed using a technique called “next-generation sequencing.” Short DNA fragments of a known sequence called adaptors are added onto the ends of the amplicon DNA molecules, creating a “library.” This library is then loaded onto a “flow cell” where sequencing takes place. This cell contains tiny wells that have small sequences of DNA attached to them that are complementary to the adapter sequences added to the amplicons. These make it so the amplicons stick to the DNA in the wells through binding. When the fragments are attached, the fragments are amplified, and thousands of copies of each fragment are generated. Another set of primers is added to the amplicon molecules, and DNA synthesis begins with multiple amplicons being sequenced at once. Each sequenced DNA amplicon will produce a “read.” After sequencing is complete, researchers can compile the different reads for all the sequences and identify mutations that are common within them.
While the introduction of DNA extraction, PCR, and next-generation sequencing as individual techniques has moved the biomedical field forward by themselves, their combined use in amplicon sequencing constitutes an approach with many advantageous applications. With uses in cancer research, pathogen detection, and disease testing, this highly sensitive and inexpensive technique allows for targeted analysis of chosen regions of the genome.
