by Fahmida Alam
Fun Rating: 4/5

Difficulty Rating: 3/5
What is the general purpose?
Viruses are obligate intracellular parasites that require host cells to survive and multiply. When viruses infect host cells, they induce various structural and functional changes, ultimately leading to cell death. The visible structural change or damage to host cells is known as a cytopathic effect (CPE). The tissue culture infectious dose 50% (TCID50) assay is one of the methods used to quantify the cytopathic effect of infectious viruses in any given laboratory virus stock or clinical specimens. It indicates the amount of virus required to infect 50% of the cells in a culture. The TCID50 Assay is also known as the cell culture infectious dose 50% assay (CCID50).
Why do we use it?
In the field of Virology, quantifying the number of infectious particles in a virus stock or experimental sample is essential for determining viral infectivity. This method is useful to quantify viruses that do not form clear plaques (Plaque Assay) or foci (Focus Forming Assay). Because this assay uses CPE as a readout, it can only be used for viruses that cause cell death as a result of infection (cytopathic viruses).
How does it work?
Step 1. Permissive cell monolayer preparation. A single layer or monolayer of cells are grown in 96-well tissue culture plates (Image 2) in appropriate cell culture conditions.
Step 2. Virus dilutions. A virus stock or experimental sample is serially diluted (10-fold dilutions). As a result, the concentration of viruses in each tube is 1/10th of the previous tube, as shown in Image 1.
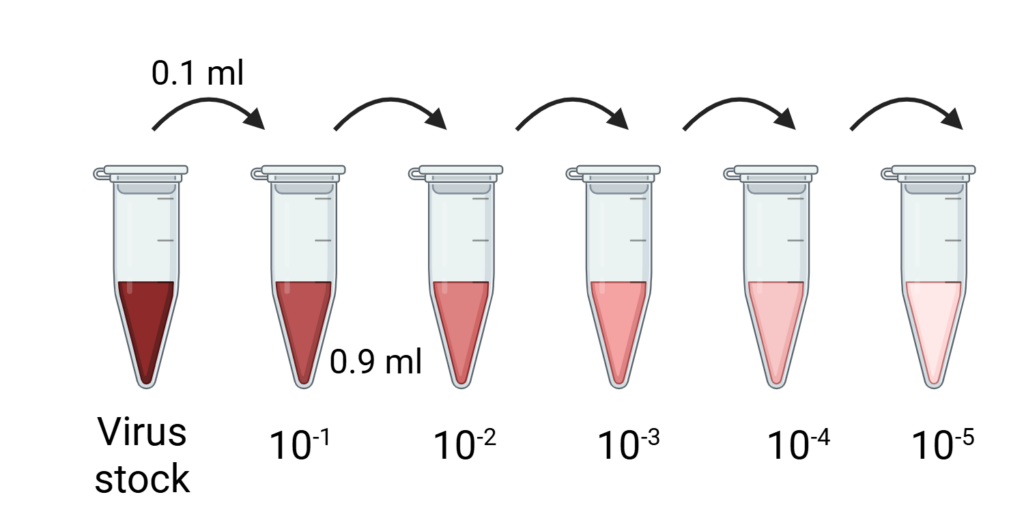
Image 1: Representative image of 10-fold serial dilution for TCID50 assay. Created by the author with BioRender.com.
Step 3. Addition of inoculum to the cell monolayer. These diluted virus samples are then added to cell monolayers. The plates are incubated at optimal temperatures (often at 33ºC or 37ºC) for 3-7 days, depending on the type of virus. During this incubation period, the virus enters the cells, multiplies within the cells, and ultimately causes CPE.
Step 4. Incubation, fixation, and staining. After the incubation period, the virus is removed from the monolayer, and the monolayer is fixed with a fixative such as formalin or formaldehyde and stained with a dye called crystal violet. The resulting plate looks like the plate shown in Image 2.
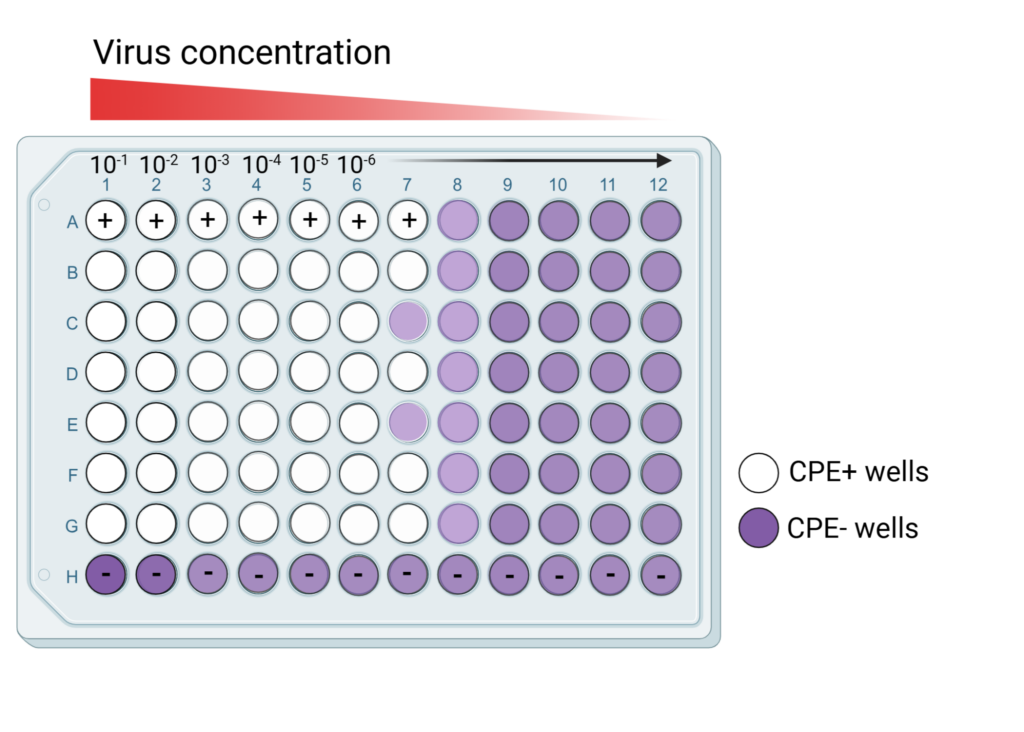
Image 2: Schematic of a TCID50 plate, ready for quantification, created by the author with BioRender.com.
Step 5. Well scoring and quantification. The crystal violet dye is meant to stain cells that were alive and still attached to the plate at the end of the incubation period. The TCID50 assay is an indirect way to quantify infectious virus particles, as it includes a probabilistic assumption in the calculation.
- For each dilution, the number of positive wells (presence of CPE/infected cells) and negative wells (no sign of CPE/infected cells) is counted. For example, in Image 2, the number of positive wells for Row A is 7 out of 12, and the number of negative wells for Row H is 12 out of 12.
- The infection rate for each dilution is calculated as follows:
Infection rate = (number of cumulative positive units or wells /
number of cumulative positive units + number of cumulative negative units) × 100
- 50% endpoint or TCID50 endpoint is the dilution that has exactly 50% infection rate. If the exact dilution is unclear from the infection rate, then it is interpolated based on the two dilutions that border 50% infection rate by calculating the proportionate distance (PD),
PD = (Infection rate above 50% – 50%) / (Infection rate above 50%) – (Infection rate below 50%)
- Log (TCID50) = log (dilution with infection rate above 50%) + (-PD) × log dilution factor.
Finally, TCID50 = 10Log (TCID50).
You can plug in numbers here to get the TCID50 of your imaginary (or real) samples!
The TCID50 assay is a useful tool for scientists to quantify the number of infectious particles present in a laboratory virus stock or clinical specimen.
